A comprehensive repository of Acinetobacter baumannii to understand the molecular landscape of antimicrobial resistance



Acinetobacter baumannii (AB) is identified as the most critical drug resistant (DR) pathogen by WHO
given that it is exhibiting increasing resistance to conventional treatments in recent years. It is
therefore imperative that DR determinants and trends of antibiotic resistance are understood across
various clinical isolates of AB. To attain this, it is required that the genomic determinants of various
clinically resistant phenotypes are consolidated along with comprehensive annotation so as to
understand the entire molecular landscape of drug resistance in AB.
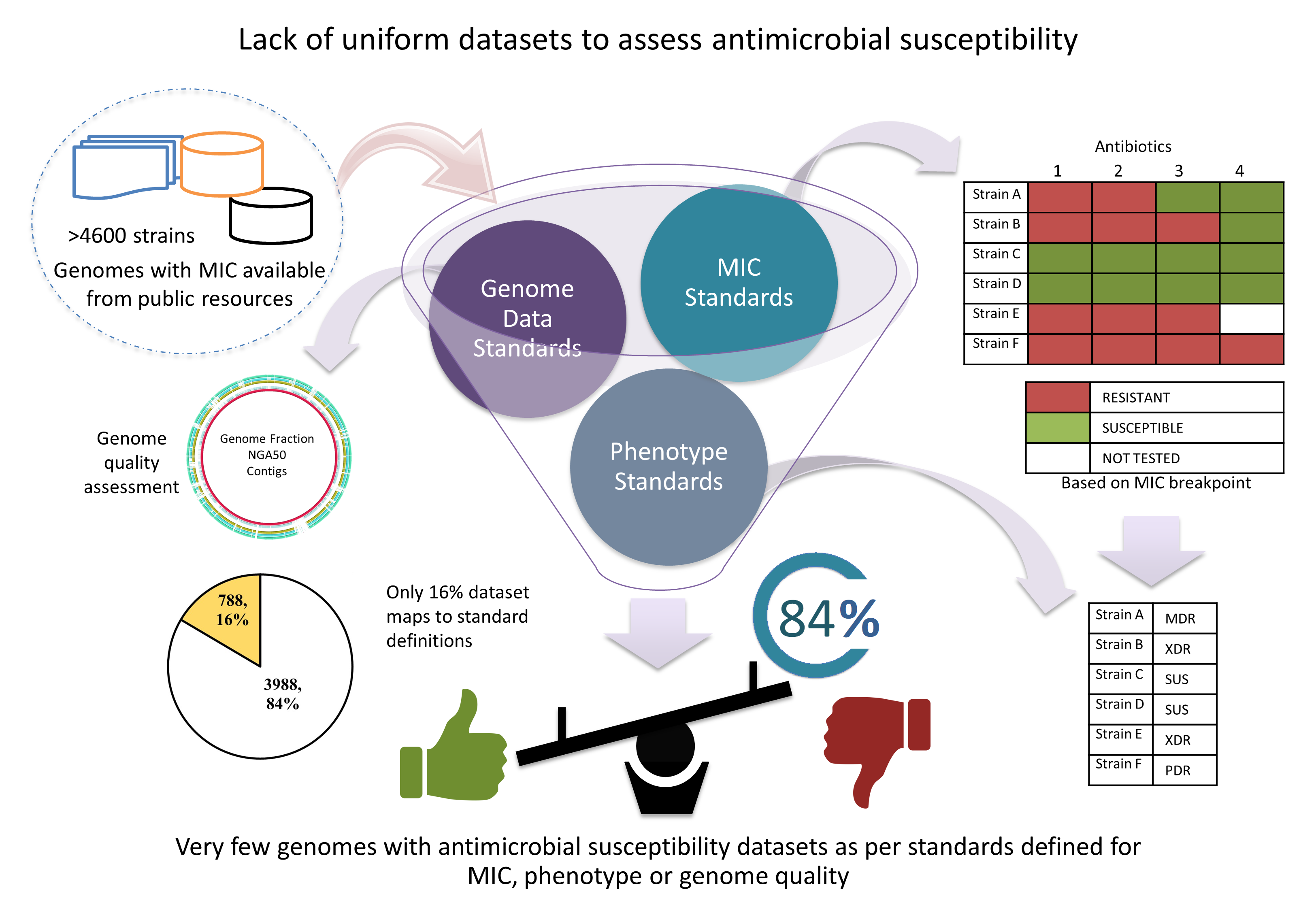
Towards this, Ab-AMR is developed with the drug resistance profile of AB genomes qualifying
CLSI/EUCAST standards, WGS standards and standards for acquired resistance profile i.e.
susceptible, multi-drug resistant (MDR), extensively-drug resistant (XDR) and pan-drug resistant
(PDR) as suggested by CDC and ECDC. DR determinants with their comprehensive annotation
including their pan-genome analysis across 788 clinical isolates have also been included in Ab-AMR. In
Ab-AMR, the reference coordinates of the revised reference genome are used to represent manually
curated and comprehensively annotated data on essential genes, genes with known pathway, PDB
structures, reported drug targets, transcription factors, sigma factors and two component systems.
Ab-AMR is made using the standard php-mysql framework and offers various search tools including a
query builder that facilitates query on over 60 different features for addressing complex questions like
core genes which are also essential and have a role to play in drug resistance with no known human
homolog, and no human octamer etc. Ab-AMR offers a centralized data resource for systematic mapping
of DR determinants, both plasmid and chromosomal mediated, along with deep annotation of clinical
isolates.
EUCAST and CLSI breakpoints for class aminoglycoside, Beta-lactams- Carbapenems, Fluoroquinolone, Beta-lactam inhibitor, Cephalosporin, Sulfonamides, Penicillins/beta-lactamase, Polymyxins, Tetracycline have been used to classify clinical isolates as susceptible, MDR, XDR and PDR. https://www.eucast.org/clinical_breakpoints.
As per EUCAST criteria, genome fraction >=75%, NGA50 >=20000 and number of contigs <=500 was used to identify good quality genomes for AST. https://www.sciencedirect.com/science/article/pii/S1198743X16305687.
According to the standard international terminologies, for AB, MDR is defined as acquired non-susceptibility to at least one agent in three or more antimicrobial categories, XDR is defined as non-susceptibility to at least one agent in all but two or fewer antimicrobial categories (i.e., bacterial isolates remain susceptible to only one or two categories) and PDR is defined as non-susceptibility to all agents in all antimicrobial categories. https://pubmed.ncbi.nlm.nih.gov/21793988/.