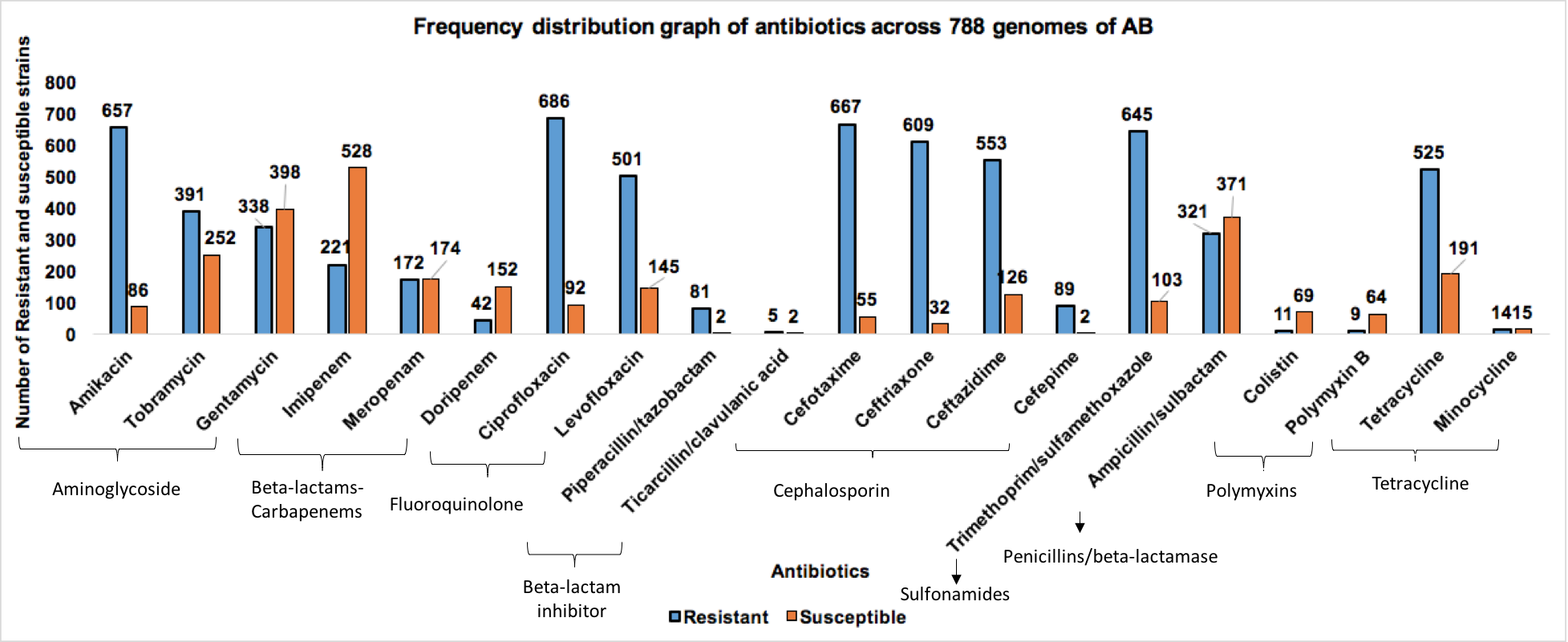
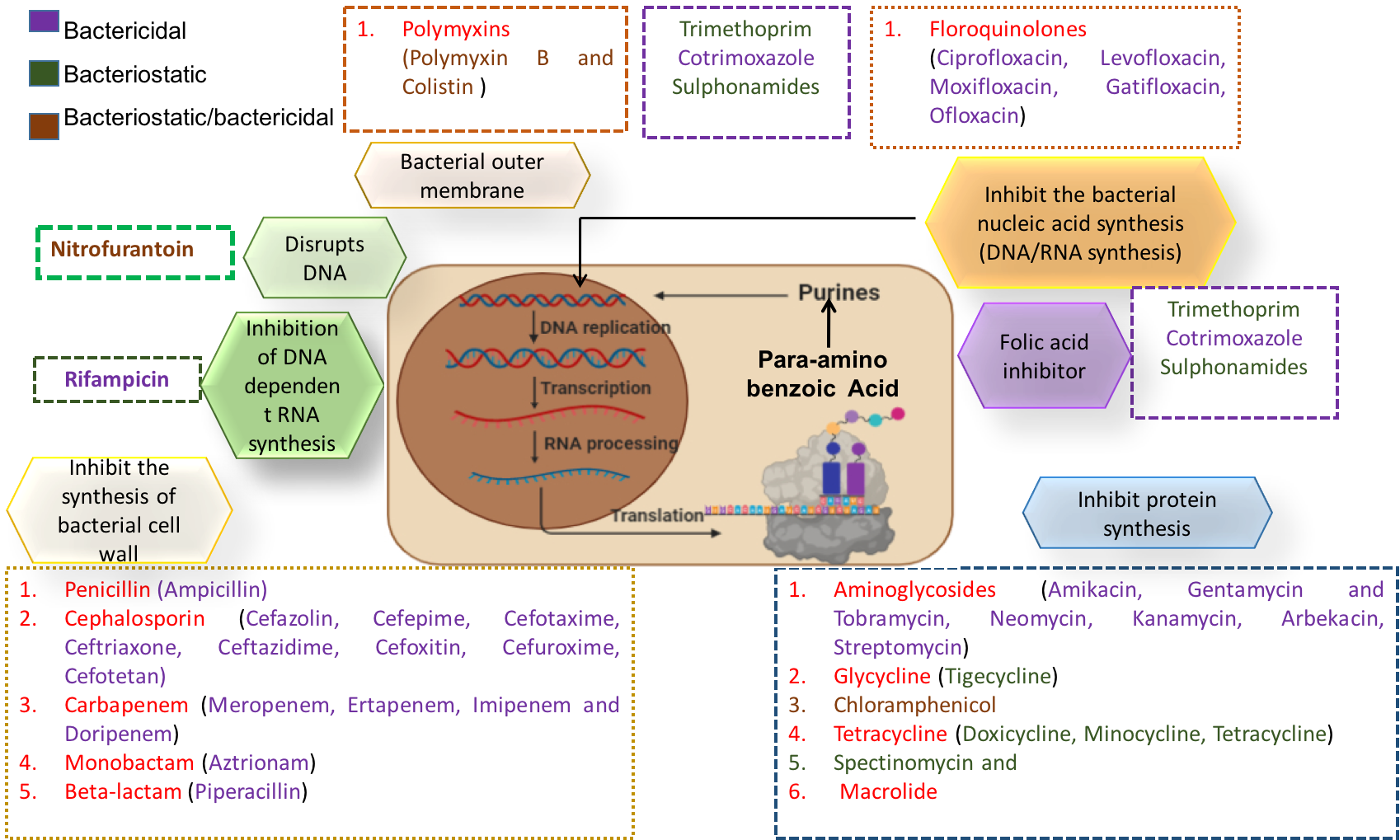
A comprehensive repository of Acinetobacter baumannii to understand the molecular landscape of antimicrobial resistance



| S.No. | Ab-AMR Features | No. of entries |
| 1 | Resistance profile | 788 genomes |
| 3 | Antibiogram (CLSI standards) | 788 clinical isolates |
| 4 | Resistant determinanats | 364 DR determinants |
| 5 | Essential genes | 614 genes |
| 6 | Drug targets | 81 genes |
| 7 | Pathways | 1334 genes |
| 7 | Reactions | 1211 |
| 8 | PDB | 221 structures |
| 9 | Transcription factors | 118 genes |
| 10 | Sigma factors | 4 genes |
| 11 | Two component system | 14 genes |
| S.No. | Resistance mechanism | No. of associated genes |
| 1 | Antibiotic inactivation | 273 |
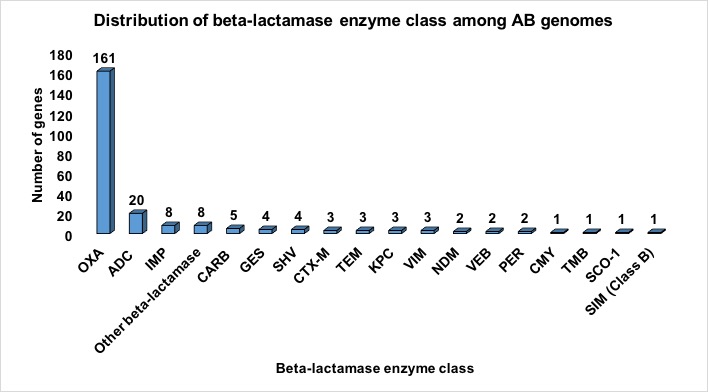
| I) | Beta-lactamases | 232 |
| II) | Aminoglycoside modification | 30 |
| III) | Chloramphenicol acetyltransferases (AACs) | 4 |
| 2 | Efflux pumps | 46 |
| 3 | Target modification | 19 |
| I) | Target replacement | 10 |
| II) | Target alteration | 7 |
| III) | Target protection | 2 |
| 4 | Other mechanisms | 14 |
| 5 | Permeability defects | 12 |